Breadcrumb
- Home
- Methodology
- Enzyme Cytochemistry
Enzyme Cytochemistry
Main navigation
Enzyme-Cytochemistry identifies an enzyme within a tissue or cell type by a specific substrate reaction resulting in a visible marker.
Substrate - the substance acted upon by an enzyme.
Trapping agent - binds to the substrate and contains a visible reaction product such as a heavy metal or a colored reaction product.
Enzymes may be classified by their method of demonstration.
A. Simultaneous capture describes the procedure where a reagent in the incubation medium combines with the reaction product. An example of this technique is the diazo method for alkaline phosphatase.
B. Post-incubation coupling is based on the production of an insoluble reaction product which is then coupled with a colored or opaque substance. Rutenburg and Seligman for acid phosphatase is an example.
C. Self-colored substrate reaction is obtained when a water-soluble dye is made insoluble due to the removal of a hydrophilic group, by the enzyme. A colored precipitate is then produced at the site of enzyme activity.
D. Intramolecular rearrangement produces a colored insoluble participate at the sites of enzyme activity of an otherwise colorless substrate.
Enzymatic reactions are often coupled with immunochemical methods.
Immunochemical Methods
Immunocytochemistry or immunohistochemistry is the identification of a tissue constituent (protein, lipopolysaccharide, etc.) in situ by means of a specific antigen/antibody reaction tagged by a visible label. immunohistochemistry usually refers to a light level detection and immunocytochemistry usually refers to detection at EM resolution.
The antigen is the protein or target of interest. An antibody is a structure created by the immune system to seek out and bind with a specific antigen. An antigen is composed of many segments called epitopes. The epitope is the region that will bind with the antibody.
Because of the special relationship between the antigen and antibody, care must be taken to assure that neither is altered beyond recognition by it’s counterpart, while at the same time maintaining the integrity of the tissue or cell in which the antigen resides.
Antibody structure
The overall structure of the antibody is a Y shape. The combining (or reactive) sites are located on the ends of the two arms. The structure of these two ends vary from one antibody to another. The third region is not involved in antigen binding. The reactive sites of the antibody are formed by four polypeptide chains bonded by bridges between sulfur-containing amino acids. A pair of identical light chains is linked to a pair of heavy chains.
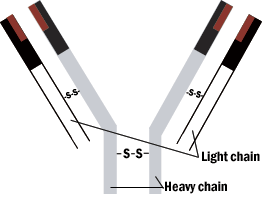
Figure 1: Schematic model of immunoglobin molecule.
Antibody structures are complex multichain proteins that have the same overall shape, yet each has unique regions that promotes binding to one antigen and not another. Folding of the antibody chains create clefts and hollows, which conform to the shape of the antigen, so that weak intermolecular forces can bind the antigen to the antibody.
Types of Antibodies:
A. Monoclonals are antibodies specific for only one antigenic site.
B. Polyclonals are a mixture of antibodies, each specific for particular epitope of the same antigen.
Types of Immuno Labeling:
A. The direct labeling method requires only one step in which a specific antibody is complexed directly to the visible marker.
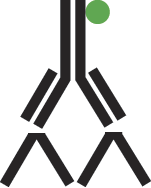
Figure 2: Schematic representation of direct labeling.
B. Indirect labeling is a two-step method which uses an antibody specific for the antigen and a secondary antibody coupled to a visible marker.

Figure 3: Indirect labeling schematic.
C. Multiple labeling (also referred to as double or triple labeling, etc.) refers to the simultaneous localization of two or more different antibodies. Obviously a different marker for each antibody must be employed.
Specific Markers:
A. Fluorescent probes may be used for obtaining antigen localization at the light level. With the invention of the laser scanning confocal microscope, this technique is especially powerful, and relatively simple.
B. Gold particles are currently the most popular tags for electron microscopy immunostaining and are used with increasing frequency at the light level as well. The particles are discreet in the electron microscope and hard to mistake. They come in a variety of sizes for double labeling. At the light microscope, a silver enhancement technique must be employed for visualization. An epi-polarizing filter enhances detection of the heavy metal label.
C. Horseradish Peroxidase (HRP)is an enzyme tag attached to an immunoglobulin with the intent of using the catalytic properties of the enzyme to form an insoluble reaction product.
D. Other enzyme markers are alkaline phosphatase, beta galactosidase, and glucose oxides. These enzymes, like HRP, require a chromogen (usually diaminobenzidine,DAB or 3-amino-9-ethylcarbazole, AEC).
Enhancement Mechanisms:
A. Hybrid binding is achieved by constructing the IgG molecule from two different antibody Fab portions, one directed against the antigen and one against the tag.
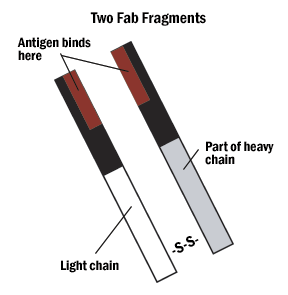
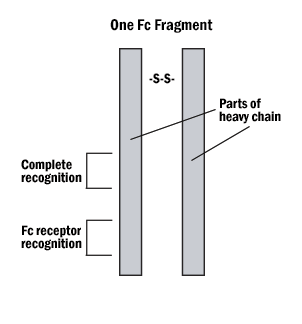
Figure 4: The Fab and Fc fragments of the antibody can be enzymatically separated for hybrid binding or very specific indirect labeling.
B. Protein A or G can be tagged on the Fc (non-reactive portion) of the IgG, leaving both reactive sites free to bind with the antigen.
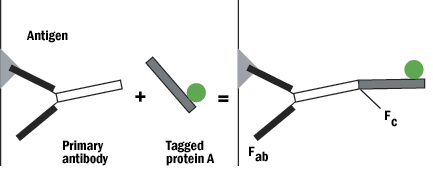
Figure 5: Protein A schematic.
C. Silver enhancement may be done when a gold particle is too small to be seen, as in a preembedment procedure, or light level observation. The silver will enucleate the gold particle.
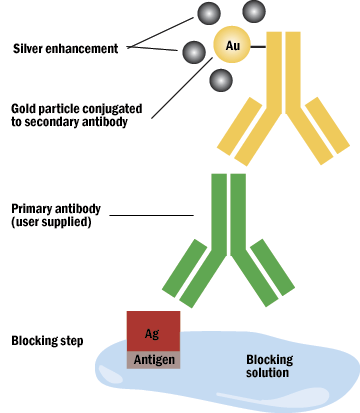
Figure 6: Silver enchancement of gold.
D. Tyramide Signal Amplification
- Boosts signal 1000x
- Easily incorporated into protocol
- Signal to noise ratio
- Sequential multi-color detection
- EM applications
- Tyramide can't diffuse
- Can use with in situ PCR or as alternative to PCR
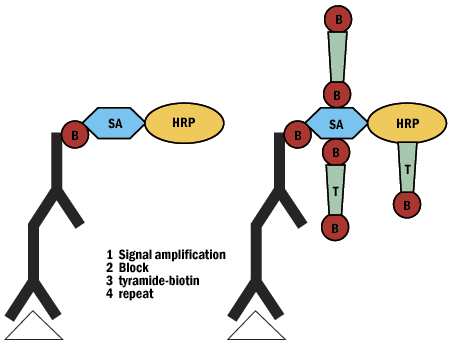
Figure 7: Tyramide signal amplification schematic.
Enzyme Controls:
A. A positive control should be employed whenever possible in parallel with the test sections to demonstrate reagent viability.
B. A negative control can be easily made by destroying the enzyme activity in a positive control by immersing in boiling water for 15 minutes, or by a specific chemical method for the enzyme to be demonstrated.
C. A useful control is to incubate one of the test sections in the reaction mixture without the substrate for the same time that the test samples are being run. Both sections are taken through the rest of the technique together.
Immuno Controls:
All experiments should be performed with a series of controls conducted simultaneously. Any “positive” result should be looked at skeptically until all controls are analyzed.
A. Adsorption. The primary antibody is reacted with an excess of antigen. This control shows that the antigen and not some other substance is responsible for the localization seen. The absorbed solution should not react with the antigen in localization protocols.
B. Tag or Unlabeled antibody. These are used separately in place of the conjugated antibody. This control determines that the specific properties of the labeled antibody are responsible for the localization.
C. Omission of primary or secondary antibodies. If labeling is seen under these conditions, then it is not the labeling intended as a result of antibody/antigen interaction.
D. Use of pre-immune sera. Collection of sera prior to production of primary antisera used in place of the primary antibody will test “positive” if some component in the immune sera other than the specific IgG is responsible for the binding. Usually it will not be possible to obtain pre-immune serum from the same animal, but it is possible to employ normal serum from the same species.
Specimen Preparation
For Enzymes
Most enzyme activity requires tissues to be as fresh as possible. Fixatives, if they can be used, should be refrigerator cold to preserve as much enzymatic activity as possible.
Some common reactions that cannot survive fixation are acid and alkaline phosphatases and esterases. Enzymes do not survive at temperatures over 55 degrees Celcius.
For Immunoreactions Pre-embedding vs. Post-embedding
A. Pre-embedding problems
1. Cell surface labeling prior to fixation
a. Frozen sections
b. Vibratome sections
2. Permeabalization may be necessary for penetration, but usually results in poor morphology.
a. Freeze/thaw
b. Detergents (Saponin, Triton-X)
c. DMSO
Pre-embedding Immunolabeling
| 1. Time required | 2-3 days |
| 2. Sectioning | Easy |
| 3. Labeling | Possible throughout section |
| 4. Serial sections | Easy |
| 5. Storage | Blocks and sections easy |
| 6. Equipment | Common in EM lab |
| 7. Disadvantages | Loss of structure, translocation with permeabalization |
B. Post-embedding immunolabeling enables more control of morphology
1. Fixation allows storage
2. Embedment media may mask antigens
3. Fixation changes protein structures
a. Etching or unmasking may be necessary
b. Fixation choices for Immunoreactions
The goal of fixation is to stabilize the cellular constituent in their native positions.
1. Chemical fixation will insure good morphology, but may compromise antigenicity and/or enzymatic activity
a. A dot immunoreactive test may help to determine antigenicity (Appendix B)
b. Overfixed tissues may respond to epitope retrieval methods (Appendix E)
2. Physical fixation may preserve antigen integrity but at the expense of morphology. For many enzymatic techniques frozen specimens may be necessary.
a. To minimize ice crystal damage a cryo protectant is used.
- sucrose
- DMSO
- glycerol
- embedment media
C. Other conditions for optimizing reactions
- pH
- Concentration of antibody and/or substrate
- Concentration of trapping reagent
- Temperature (enzymes usually will not survive to temperature over 55 degrees celcius)
- Reaction or fixation times
D. Blocking reagents can be used to block background staining due to cross-reactivity or multispecificity.
Closely related antigens whose molecular shape and charge distribution may be similar to the original antigen and may fit the same antibody. They are said to cross-react.
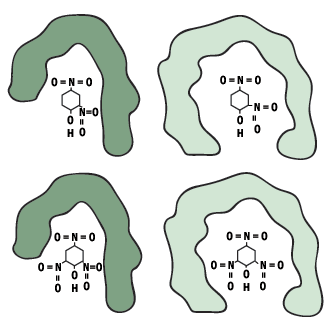
Figure 8: Two antibodies cross react with a related antigen (DNP)
Very different antigens have been known also to bind with a single antibody, the close contacts of the edges of the antigen and the amino acid sites that form the walls of the combining site of the antibody may cause it to become bound. This feature is termed multispecificity.
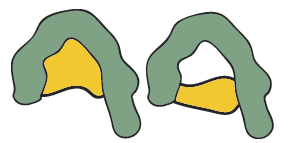
Figure 9: The same antibody reacting with two very different antigens (multispecificity).